Molecular surface
The MolecularSurfaceSsettings object controls various settings related with molecular surface. Here the molecular surface includes:
Solvent accessible surface (SAS)
van der Waals surface, a special case of SAS with probe radius equal to 0)
Solvent-excluded surface (SES) or Connolly surface
Solvent accessible surface
Here we show a example of draw SAS for the protein kras.
>>> from ase.io import read
>>> from batoms import Batoms
>>> kras = read("kras.pdb")
>>> kras = Batoms("kras", from_ase = kras)
>>> kras.molecular_surface.draw()
>>> kras.get_image(padding = 3, output = "ms_sas_kras.png")
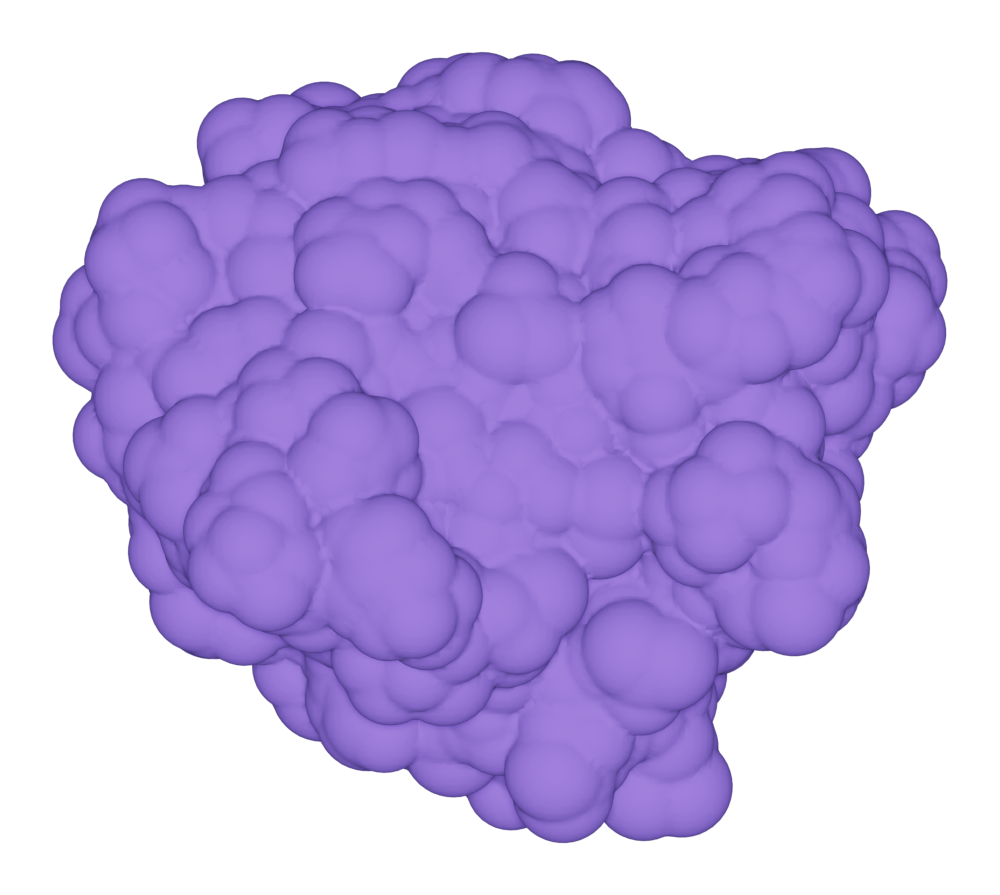
You can print the default setting by:
>>> kras.molecular_surface.settings
name select probe resolution color
1 all 1.400 0.500 [0.0 1.0 1.0 1.0]
The default probe radius is set to be 1.4. One can change it by:
>>> kras.molecular_surface.settings["1"].probe = 1.2
You can get the solvent accessible surface area (SASA) by:
>>> area = kras.molecular_surface.settings.get_sasa("1")
Area: 7977.796, Volume: 33748.846
Analytical SAS area calculated by MSMS is 8119 Ų.
You can get the solvent accessible surface with respect to each atoms by:
>>> area = kras.molecular_surface.settings.get_psasa()
Solvent-excluded surface
Here we show a example of draw SES for the protein kras.
>>> from ase.io import read
>>> from batoms import Batoms
>>> kras = read("kras.pdb")
>>> kras = Batoms("kras", from_ase = kras)
>>> kras.molecular_surface.settings["1"].type = "SES"
>>> kras.molecular_surface.draw()
>>> kras.get_image(padding = 3, output = "ms_ses_kras.png")
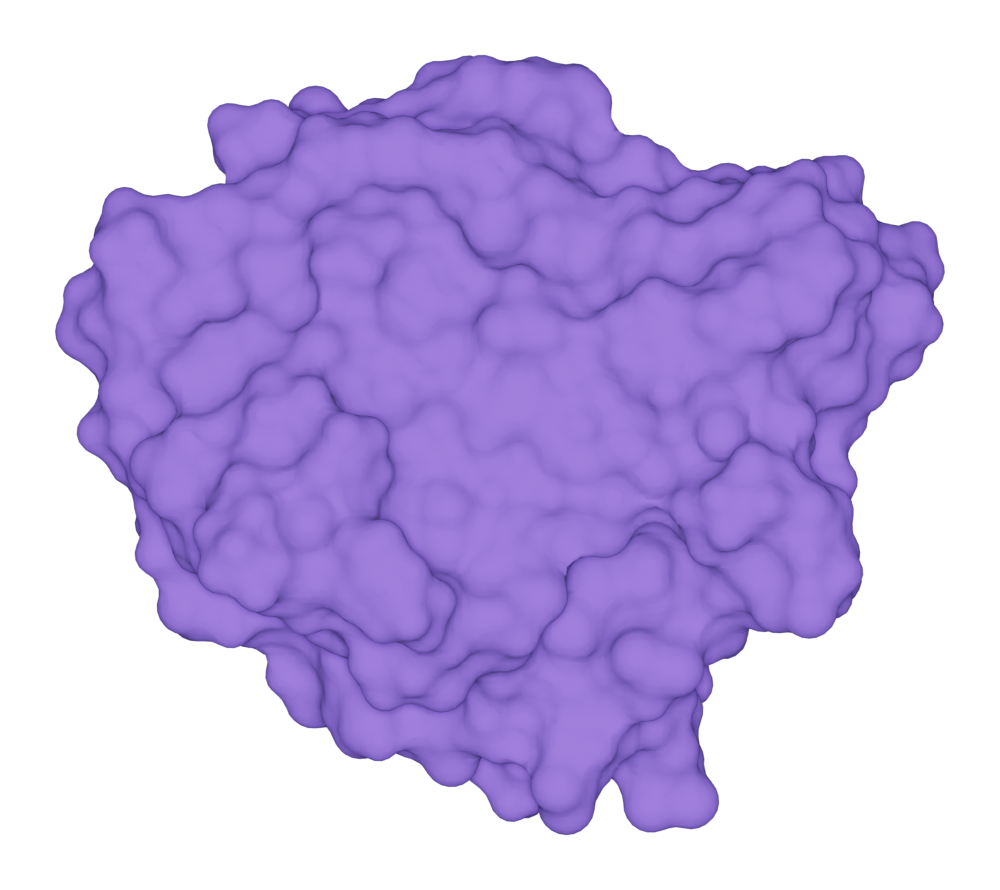
You can get the solvent-excluded surface area (SESA) by:
>>> area = kras.molecular_surface.settings.get_sesa("1")
SES: area: 7071.490, volume: 23277.100
Analytical SES area calculated by MSMS is 7080 Ų.
Note
0.4 is usually a reasonable resolution to get accurate SAS area and SES area close to analytical results.
Real-time rendering of SAS
Metaball method is used to for real-time rendering of SAS.
Please read Metaball page to setup metaball.